 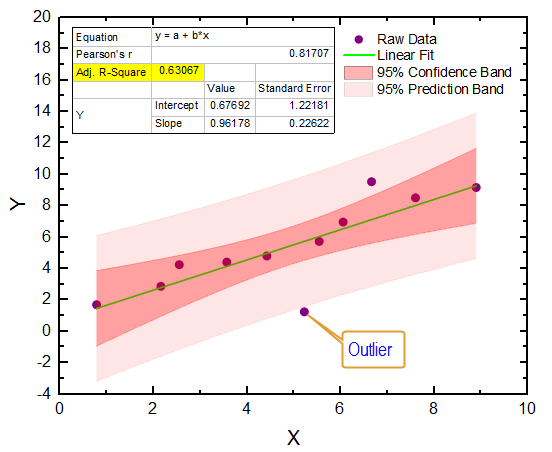  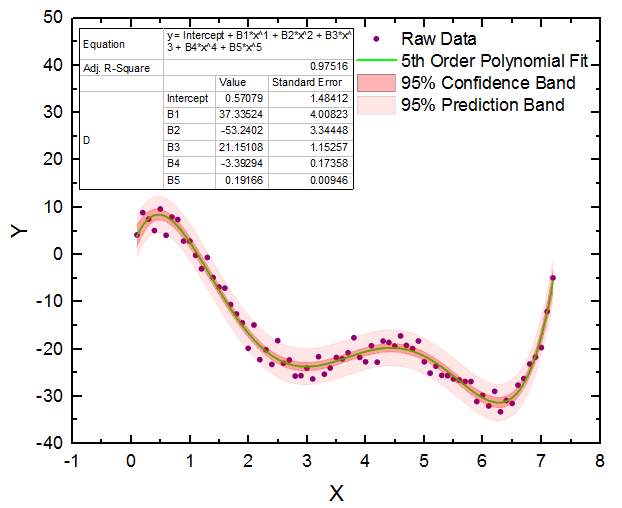
At the same time, these tutorials are well-suited to familiarize students with the principal concepts of data reduction, data transformation, error propagation, and NLLS analysis. The versatility and utility of Fit-o-mat are demonstrated below by way of four examples representative of undergraduate and graduate curricula in chemistry and the life sciences. Moreover, members of my research group routinely evaluate their experimental data and prepare figures for publication with the software. level, both in the laboratory and for homework assignments. Among other use cases, the program has found deployment in a course on Synthetic Biology at the M.Sc. Over the course of several months, Fit-o-mat has been applied in both teaching and research settings and has thereby proven to be a versatile and dependable resource for quantitative data evaluation and presentation. Tutorial mode to facilitate deployment in the classroom (see Figure 1B) Program session files to promote ready data exchange Style sheets for preparing consistent figures Publication-quality graphics in various vectorized and bitmapped output formats NLLS fitting to arbitrary, user-defined Python functions, including discontinuous and numerical functions Options for data reduction and transformation Released under the GNU General Public License (version 3.0 or later)ĭata import from text and Microsoft Excel files Fit-o-mat offers the following key features, certain of which are treated in more detail in the next section:Ĭross-platform, Python 3-based architecture The program and tutorial files are also included in the Supporting Information. (16) Version 0.510 of the program source code is available free of charge from or from an accompanying manual covers program requirements, installation, and use. (10)įit-o-mat features a graphical interface based on the PyQt5 (18) framework and thereby provides a convenient front end for numerical procedures for NLLS analysis implemented in the Python libraries NumPy (14) and SciPy, (15) and for data plotting implemented in matplotlib. (4,5) In addition to Arrhenius-type problems, students, instructors, and researchers encounter nonlinear relations in many areas of chemistry and biochemistry, such as (but not limited to) enzyme kinetics, (6−8) protein folding, (9) and cyclic voltammetry. 
) is determined, commonly by using the Levenberg–Marquardt algorithm.Starting from an initial parameter set, an optimum set of parameters ( a 1, a 2, It should be noted that in eq 3, the errors σ i serve as weighting factors for the individual data points i. In this procedure, the agreement between experimental data y i and the (original, nonlinear) functional relationship f( x i) is usually evaluated as χ 2, which refers to the sum over i of the squared deviations between y i and f( x i): (3)In an iterative manner, parameter values are varied in such a way that the target function χ 2 is minimized, and the process is hence called nonlinear least-squares (NLLS) curve fitting. To reiterate, transformation is best avoided, and parameter values should be determined by nonlinear curve fitting of the original data. (Accordingly, in this example k( T) and T are the dependent and independent variables, respectively.) To determine the parameters A and E a, the rate constant of a reaction of interest is hence recorded at several temperatures. For instance, the temperature dependence of rate constants k in chemistry and biochemistry can often be modeled according to the Arrhenius equation: (1,2) (1)where A is the pre-exponential factor, E a is the activation energy, R is the gas constant, and T is the absolute temperature. ), serves to verify (or invalidate) physical models underpinning the experimental data and to determine characteristic parameters ( a 1, a 2,.Analysis according to functional relationships, denoted y = f( x, a 1, a 2, In the classroom, as in the laboratory, experimental observables, called dependent variables y, are routinely evaluated as functions of one or several independent variables, denoted as x. The quantitative analysis of data and their graphical presentation are essential across many disciplines, including chemistry and the life sciences. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
